Plot a Discrete Survival Function as a Step Function
Source:R/geom_survival_discrete.R
geom_survival_discrete.Rdgeom_survival_discrete() renders the discrete survival function
\(S(x) = 1 - F(x)\) as a right-continuous step function with horizontal
segments, dashed vertical jumps, open circles at the lower limit of each
jump, and closed circles at the upper limit.
Usage
geom_survival_discrete(
mapping = NULL,
data = NULL,
stat = StatSurvivalDiscrete,
position = "identity",
...,
na.rm = FALSE,
show.legend = NA,
inherit.aes = FALSE,
fun = NULL,
cdf_fun = NULL,
pmf_fun = NULL,
xlim = NULL,
support = NULL,
args = list(),
open_fill = NULL,
vert_type = "dashed",
show_points = NULL,
show_vert = NULL
)
StatSurvivalDiscrete
GeomSurvivalDiscreteFormat
An object of class StatSurvivalDiscrete (inherits from Stat, ggproto, gg) of length 3.
An object of class GeomSurvivalDiscrete (inherits from Geom, ggproto, gg) of length 5.
Arguments
- mapping
Set of aesthetic mappings created by
aes(). If specified andinherit.aes = TRUE(the default), it is combined with the default mapping at the top level of the plot. You must supplymappingif there is no plot mapping.- data
The data to be displayed in this layer. There are three options:
NULL(default): the data is inherited from the plot data as specified in the call toggplot().A
data.frame, or other object, will override the plot data. All objects will be fortified to produce a data frame. Seefortify()for which variables will be created.A
functionwill be called with a single argument, the plot data. The return value must be adata.frame, and will be used as the layer data. Afunctioncan be created from aformula(e.g.~ head(.x, 10)).
- stat
The statistical transformation to use on the data for this layer. When using a
geom_*()function to construct a layer, thestatargument can be used to override the default coupling between geoms and stats. Thestatargument accepts the following:A
Statggproto subclass, for exampleStatCount.A string naming the stat. To give the stat as a string, strip the function name of the
stat_prefix. For example, to usestat_count(), give the stat as"count".For more information and other ways to specify the stat, see the layer stat documentation.
- position
A position adjustment to use on the data for this layer. This can be used in various ways, including to prevent overplotting and improving the display. The
positionargument accepts the following:The result of calling a position function, such as
position_jitter(). This method allows for passing extra arguments to the position.A string naming the position adjustment. To give the position as a string, strip the function name of the
position_prefix. For example, to useposition_jitter(), give the position as"jitter".For more information and other ways to specify the position, see the layer position documentation.
- ...
Other parameters passed on to
ggplot2::layer().- na.rm
If
FALSE, the default, missing values are removed with a warning. IfTRUE, missing values are silently removed.- show.legend
Logical. Should this layer be included in the legends?
NA, the default, includes if any aesthetics are mapped.FALSEnever includes, andTRUEalways includes. It can also be a named logical vector to finely select the aesthetics to display. To include legend keys for all levels, even when no data exists, useTRUE. IfNA, all levels are shown in legend, but unobserved levels are omitted.- inherit.aes
If
FALSE, overrides the default aesthetics, rather than combining with them. This is most useful for helper functions that define both data and aesthetics and shouldn't inherit behaviour from the default plot specification, e.g.annotation_borders().- fun
A discrete survival function evaluated directly on the integer support. Exactly one of
fun,cdf_fun, orpmf_funmust be provided.- cdf_fun
A discrete CDF function (e.g. pbinom). \(S(x) = 1 - F(x)\) is computed from this function on the integer support. Exactly one of
fun,cdf_fun, orpmf_funmust be provided.- pmf_fun
A PMF function (e.g. dbinom). The survival function is derived as \(1 - \mathrm{cumsum}(\mathrm{pmf})\). Exactly one of
fun,cdf_fun, orpmf_funmust be provided.- xlim
A numeric vector of length 2 specifying the range of integer support values.
- support
An optional integer or numeric vector giving the exact support points to evaluate. When supplied,
xlimis ignored.- args
A named list of additional arguments to pass to
fun,cdf_fun, orpmf_fun.- open_fill
Fill color for the open (hollow) endpoint circles. Defaults to
NULL, which uses the active theme's panel background color.- vert_type
Line type for the vertical jump segments. Defaults to
"dashed".- show_points
Logical. If
FALSE, suppresses all endpoint circles (open and closed). IfNULL(the default), circles are shown when there are 50 or fewer points and hidden otherwise.- show_vert
Logical. If
FALSE, suppresses the vertical jump segments. IfNULL(the default), segments are shown when there are 50 or fewer points and hidden otherwise.
Details
Supply exactly one of fun (a discrete survival function evaluated
directly), cdf_fun (a discrete CDF such as pbinom, from which
\(S(x) = 1 - F(x)\) is computed), or pmf_fun (a PMF such as dbinom,
from which the CDF is computed via cumulative summation and then
\(S(x) = 1 - F(x)\)).
Examples
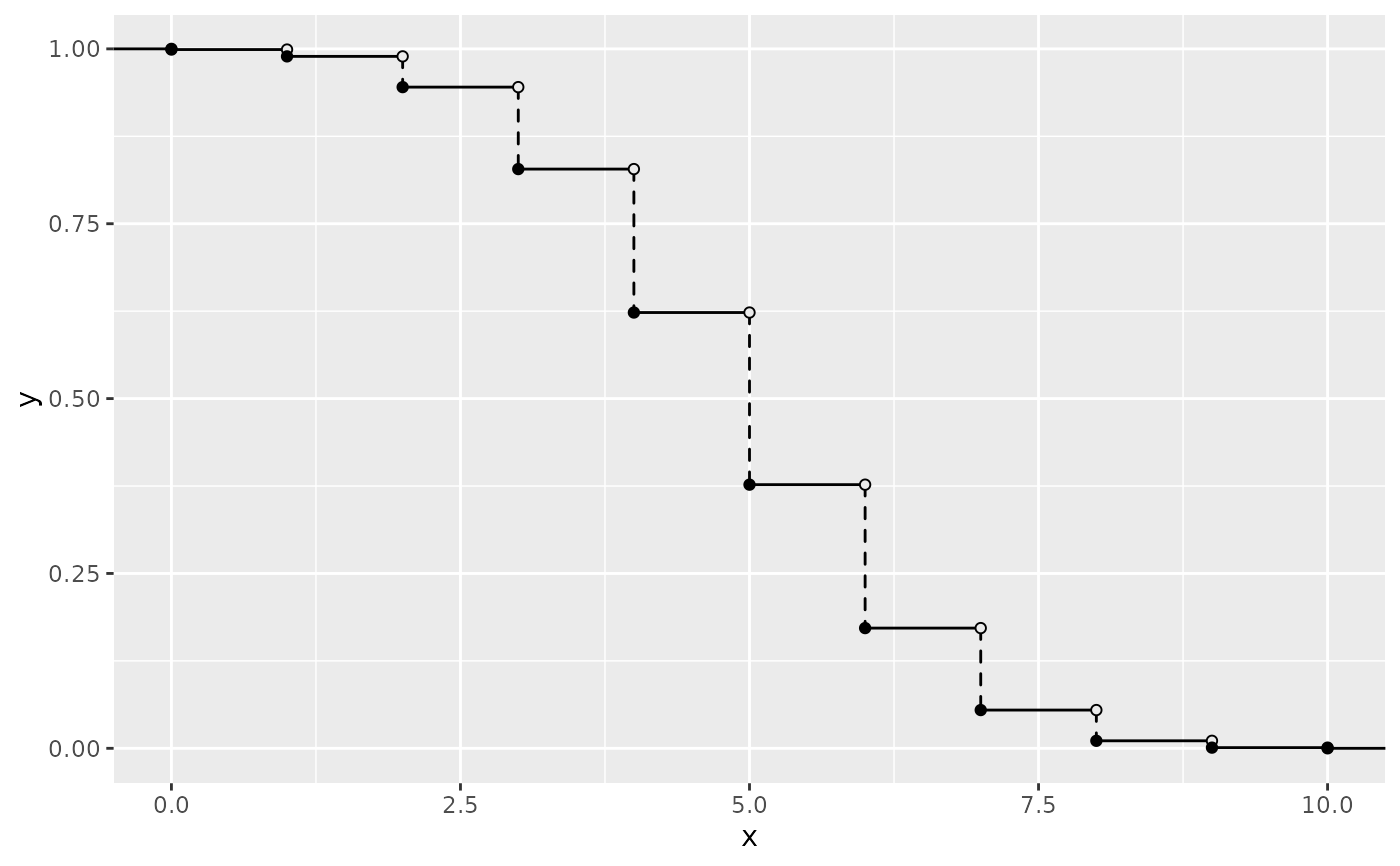
# via PMF
ggplot() +
geom_survival_discrete(pmf_fun = dbinom, xlim = c(0, 10), args = list(size = 10, prob = 0.5))
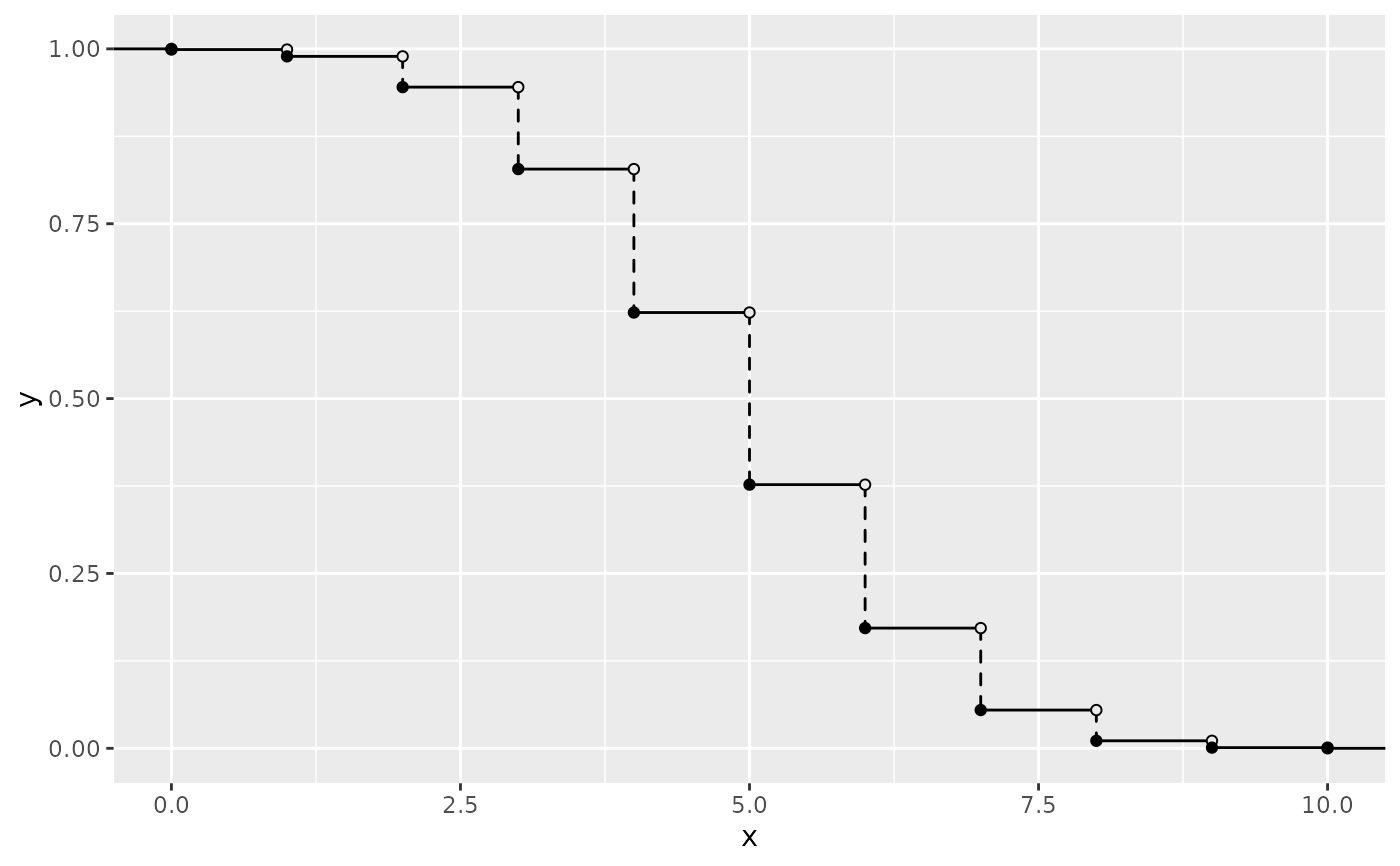
 # via CDF
ggplot() +
geom_survival_discrete(cdf_fun = pbinom, xlim = c(0, 10), args = list(size = 10, prob = 0.5))
# via CDF
ggplot() +
geom_survival_discrete(cdf_fun = pbinom, xlim = c(0, 10), args = list(size = 10, prob = 0.5))
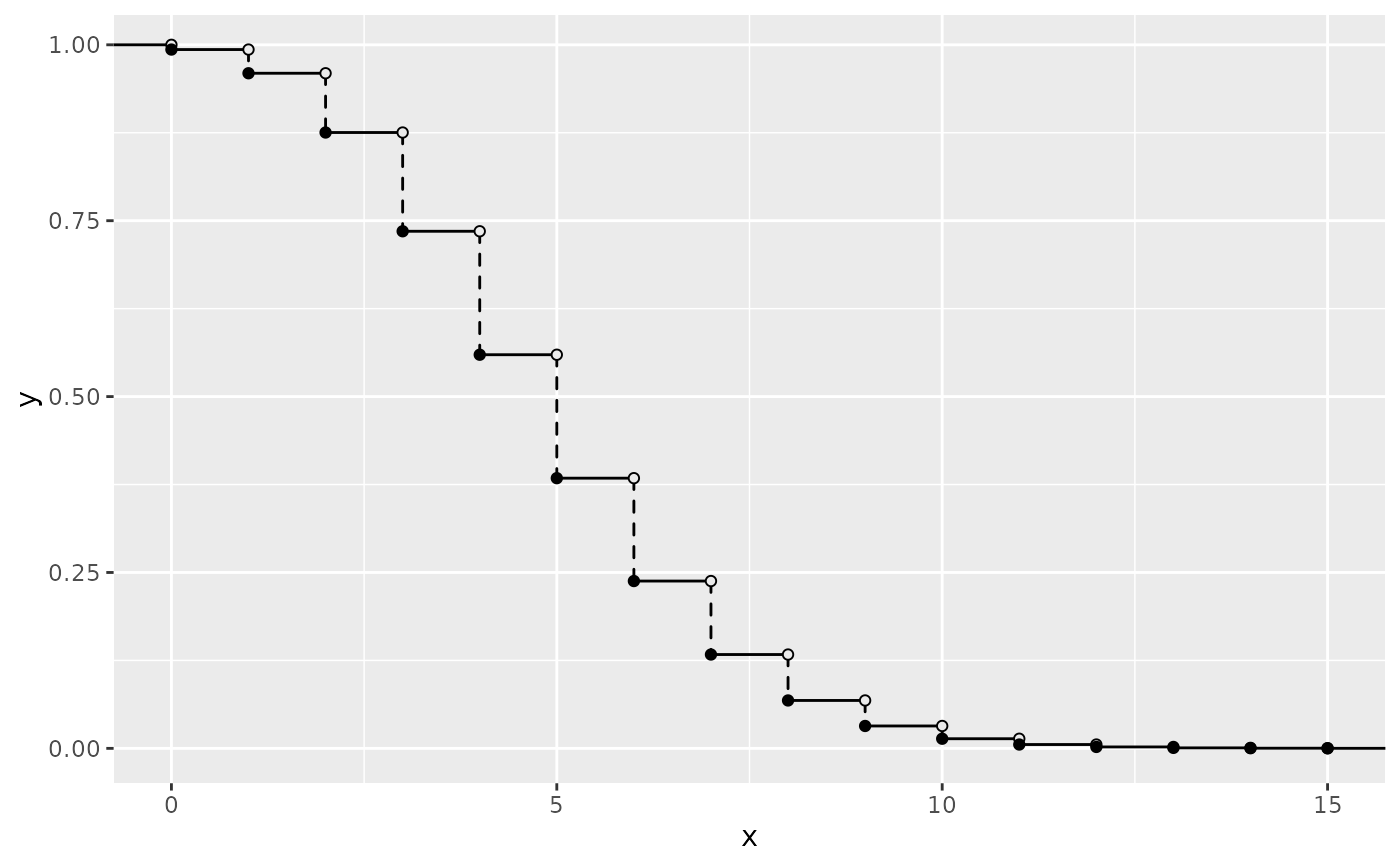
 ggplot() +
geom_survival_discrete(pmf_fun = dpois, xlim = c(0, 15), args = list(lambda = 5))
ggplot() +
geom_survival_discrete(pmf_fun = dpois, xlim = c(0, 15), args = list(lambda = 5))